SUNFLO
Pierre Casadebaig, Philippe Debaeke, Jérémie Lecoeur (INRA)
Overview
| Model category | gbCSM |
|---|---|
| Plant part | Shoot |
| Scale | Organs, Field |
| Licence | open_source |
| Operating system | Windows, Linux, MacOs |
| Programming language | Cpp, R, C++ |
| Format of model inputs and outputs | dataframe files |
| Species studied | Sunflower |
| Execution environment | Console, Application |
Scientific article
SUNFLO, a model to simulate genotype-specific performance of the sunflower crop in contrasting environmentsPierre Casadebaig,Lydie Guilioni,Jérémie Lecoeur,Angélique Christophe,Luc Champolivier,Philippe DebaekeAgricultural and Forest Meteorology, 2011 View paper
Model description
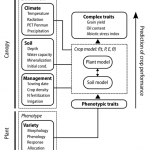
SUNFLO is a process-based model for the sunflower crop which was developed to simulate the grain yield and oil concentration as a function of time, environment (including soil, climate and management practice) and genetic diversity. In this model, the daily crop dry biomass is calculated as a function of incident photosynthetically active radiation, light interception efficiency and radiation use efficiency. The light interception efficiency is based on Beer-Lambert's law. The radiation use efficiency concept is used to represent photosynthesis at the crop scale. Broad scale processes of this framework, the dynamics of leaf area, photosynthesis, and biomass allocation to grains were split into finer processes (e.g leaf expansion and senescence, responses to environmental stresses) to reveal genotypic specificity and to allow the emergence of genotype-by-environment interactions. Globally, the SUNFLO crop model has about 50 equations and 64 parameters.
Some case studies
Some recent studies using SUNFLO:
- A model-based approach to assist variety evaluation in sunflower crop (https://doi.org/10.1016/j.eja.2016.09.001)
- Quantifying physiological determinants of genetic variation for yield potential in sunflower. SUNFLO: A model-based analysis (https://doi.org/10.1071/FP09189)
- Heliaphen, an Outdoor High-Throughput Phenotyping Platform for Genetic Studies and Crop Modeling (https://doi.org/10.3389/fpls.2018.01908)